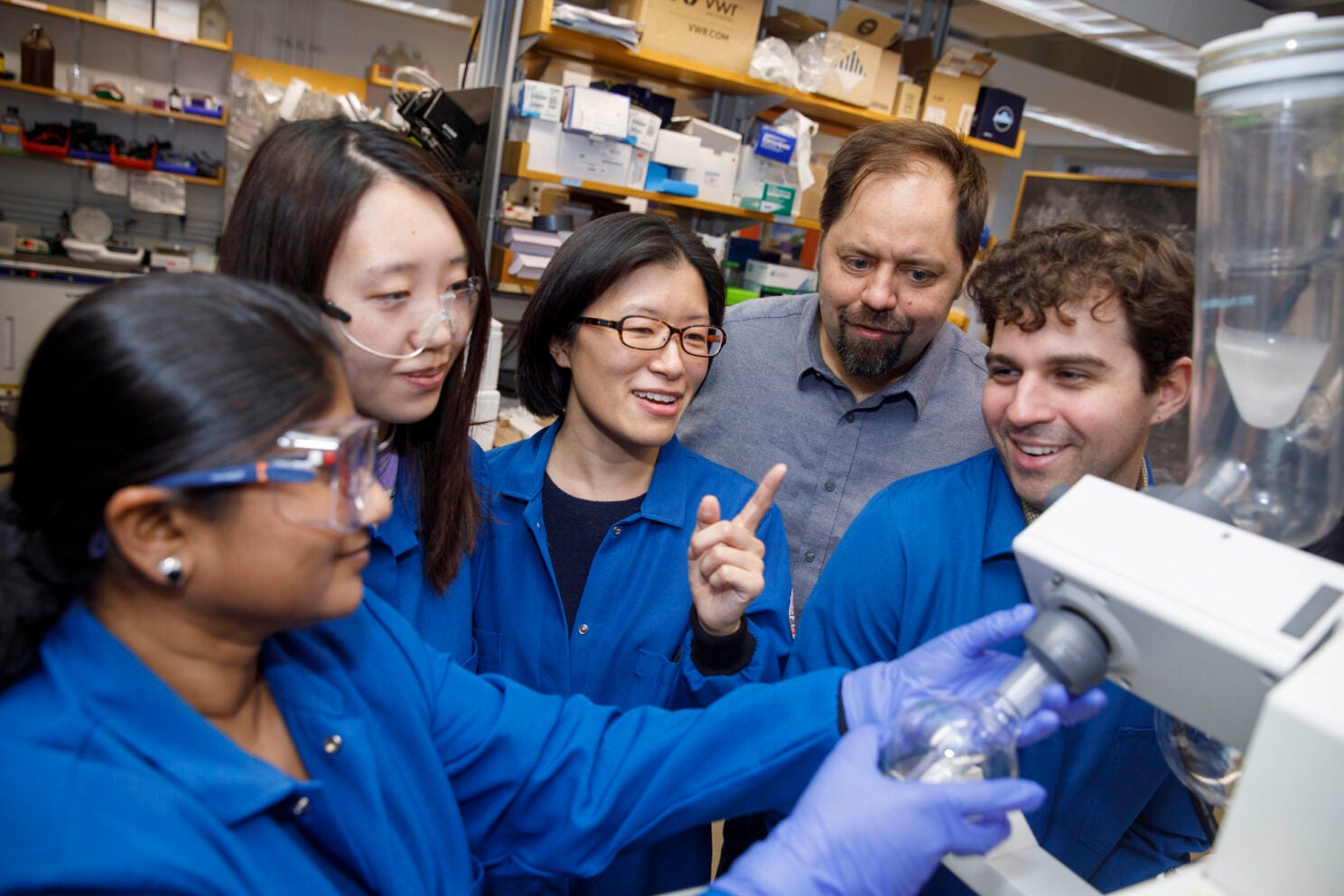
Cristina Woo (center) in her lab with study co-authors Nandini Vallavoju (from left), Wenqing Xu, Ralph Mazitschek, and Connor Payne.
Kris Snibbe/Harvard Staff Photographer
In the battle against cancer and other diseases, scientists are developing molecular weapons that can be used to stop uncontrollable cell growth.
A team of Harvard and Massachusetts General Hospital scientists have found that “cyclimids,” a class of binding molecules known as ligands, offer a promising and efficient approach to removing disease-causing or malfunctioning proteins. Their distinct properties enable scientists to attack errant proteins at their molecular roots.
“For over a year we had been tackling the question of what are the natural ligands that are recognized by cereblon, a protein crucial to targeted degradation,” said senior co-author Christina Woo, Morris Kahn Associate Professor of Chemistry and Chemical Biology. “This study comprehensively characterizes these ligands to provide new insights to cereblon biology and how to hijack it.”
In recent years, scientists have engineered small molecules to specifically target proteins associated with disease. These molecules have two roles: they latch onto the target protein that needs to be removed, and their “warhead” engages with part of the cellular cleanup system, often binding to a protein called cereblon. Together, these specialized molecules form what scientists call a ternary complex. Once this complex is established, the target protein gets effectively marked for disposal by the cell’s proteasome, which acts like a cellular recycling system. The success of this process — removing specific proteins — is dependent on the design and efficiency of the molecular warhead, making them crucial elements in the development of therapies for various diseases, including cancer.
In the researchers’ paper published in Cell: Chemical Biology, they found that minor structural changes on the cereblon ligand can dramatically alter biological activities in cells. In collaboration with the Mazitschek Lab, which has done extensive research into the identification of disease-relevant molecular targets, the researchers introduced a systematic biochemical approach for quantifying ternary complex formation. This method allows researchers to predict the cellular degradation activity of cyclimids more effectively, streamlining the development process.
“We have given the community a powerful and affordable microscope with our method,” Ralph Mazitschek, co-senior author, said. “We have established a comprehensive, reliable, robust, and sensitive profiling platform that is applicable to virtually any of these small molecule degraders and molecular glue degraders.”
“This was a collaboration in the truest sense of the word,” said Connor Payne, postdoctoral fellow in Mazitschek’s lab. “We had different expertises, and different technologies that we were developing so the synergy between them was really, really beautiful to see come to fruition.”
Going forward, Woo and Mazitschek are optimistic that cyclimids and their screening platforms will be incorporated into protein-degradation strategies, which could be useful in developing drugs and treating cancers.
“I think our research will ultimately facilitate profiling many more molecules against desired targets and arrive at more selective and efficacious molecules faster,” Woo said. “There are a lot of different directions this could take us.”
The following graduate students in Woo’s lab also took part in the research: Saki Ichikawa, Wenqing Xu, Chia-Fu Chang, Nandini Vallavoju, Hope A. Flaxman, and Spencer Frome.